LonGenEx (DALIA)
LonGenEx is an adapted version of CoPHI, tailored to the exploration of gene expression data from vaccine trials. It was originally developped using the DALIA-1 trial data for an HIV vaccine.
Try out the online tool here.
Dataset details
The online tool comes with 4 preloaded datasets corresponding to two vaccine trials:
- DALIA-1, HIV vaccine trial (original data)
- rVSV, ebola vaccine trial (original data)
In these trials, gene expression levels are measured for a number of patients at different points in time. In the preloaded datasets, the original data was normalized and flatten by aggregating (median) patients and probe data for each gene. Each dataset has two versions, with one holding ‘variation’ data.
Datasets are extended by associating to each gene a Chaussabel’s modules V2. For now, the Chaussabel modules are slightly transformed such that one gene solely belongs to one module. Gene originally belonging to several modules are associated to the Mx.y module that is the smallest relative to x.
Associated publications
The following publication is associated to the DALIA-1 data:
Hejblum BP, Skinner J, Thiébaut R (2015) Time-Course Gene Set Analysis for Longitudinal Gene Expression Data. PLoS Comput Biol 11(6): e1004310.
The following publication is associated to the rVSV-ZEBOV data:
Rechtien A, et al. (2017) Systems Vaccinology Identifies an Early Innate Immune Signature as a Correlate of Antibody Responses to the Ebola Vaccine rVSV-ZEBOV. Cell Reports. 2017 Aug;20(9):2251-2261.
The gene modules come from the following publication:
Chaussabel, D, Baldwin, N (2014) Democratizing systems immunology with modular transcriptional repertoire analyses. Nat Rev Immunol 14, 271–280.
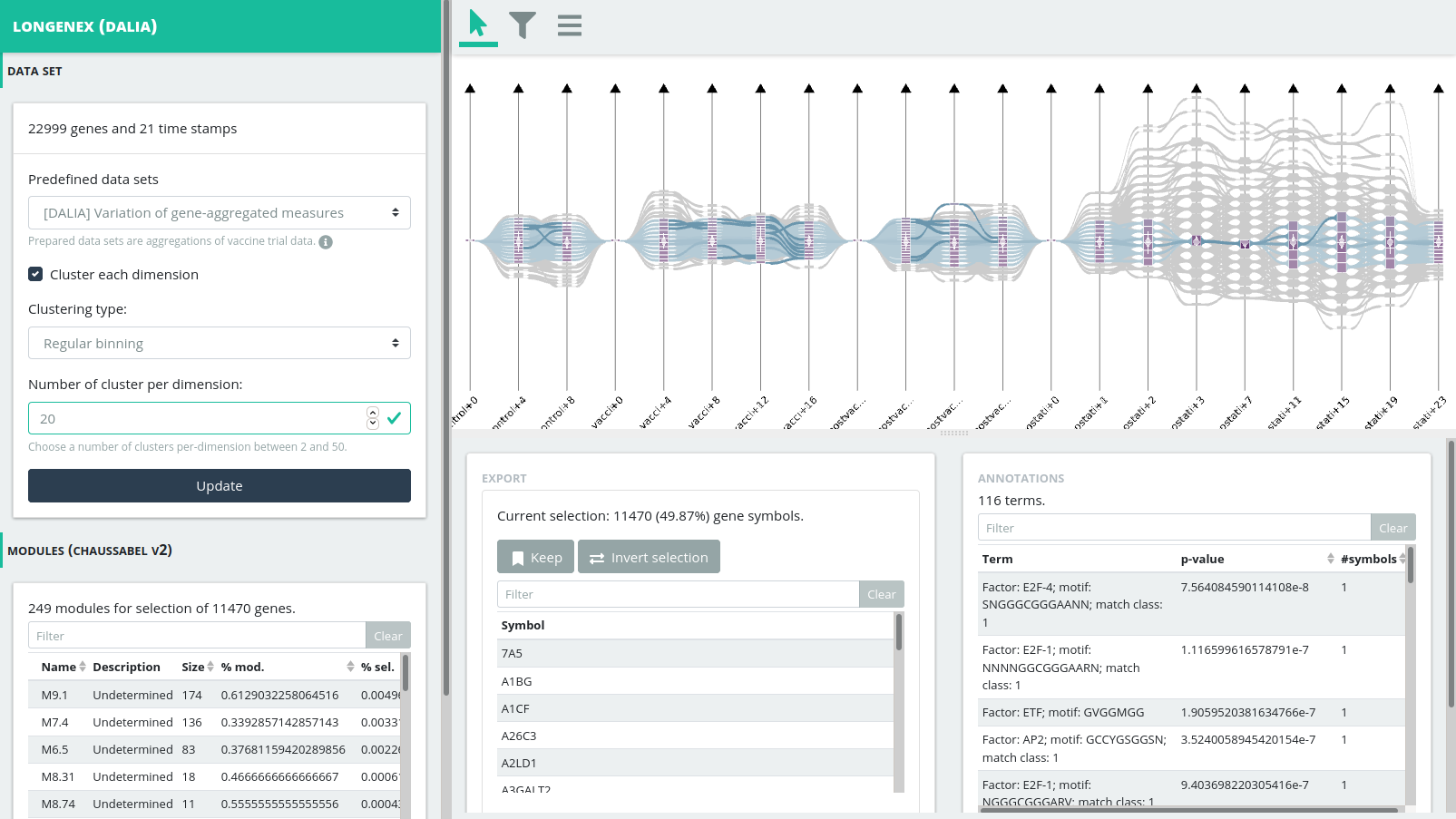
Interacting with the tool
Selecting visual elements in the visual representation corresponds to selecting subsets of genes. The tool provides side views with detail on the selected subset of genes.
- Annotation (bottom right): The annotation view shows annotation for the gene subset as queried on GProfile;
- Module (bottom left): The module view updates the Chaussabel’s modules V2 list to show which module are related to the selected gene subset;
- Export (bottom): The export view allows to save multiple selections of gene subset and export them as a CSV file.
Collaborators
- David Auber
- Romain Bourqui
- Boris Hejblum
- Rodolphe Thiébaut
- …